We’re excited to share the publication of alphaDIA in Nature Biotechnology. alphaDIA is a modern, open-source search engine that brings deep learning directly to raw mass spectrometry data for data-independent acquisition (DIA) proteomics.

Figure: Wallmann et al, 2025, Nature Biotech.
A search engine for modern DIA proteomics
Mass spectrometry-based proteomics continues to generate increasingly complex datasets, especially with new, high-speed time-of-flight (TOF) detectors and advanced acquisition modes. alphaDIA addresses this complexity head-on through a feature-free processing approach, applying machine learning directly on the raw signal instead of relying on peak detection or data reduction. This enables robust analysis even for highly noisy TOF data, ensuring that all information contributes to peptide identification.
A major innovation introduced with alphaDIA is DIA transfer learning, built on the deep learning framework alphapeptdeep. By continuously fine-tuning neural networks to reflect the specific instrument and experimental setup, alphaDIA adapts to new conditions, e.g. different post-translational modifications (PTMs), without requiring specialized retraining. This flexibility allows the analysis of previously inaccessible peptide classes and significantly improves both identification depth and FDR control.
In benchmarking studies, alphaDIA matches or exceeds the performance of established tools, identifying over 120,000 precursors and nearly 10,000 protein groups in 21-minute gradients on the Orbitrap Astral. The framework supports both empirical and fully predicted libraries and integrates seamlessly with the broader alphapept ecosystem, including alphaRaw, alphaTims, directLFQ, and alphapeptdeep.
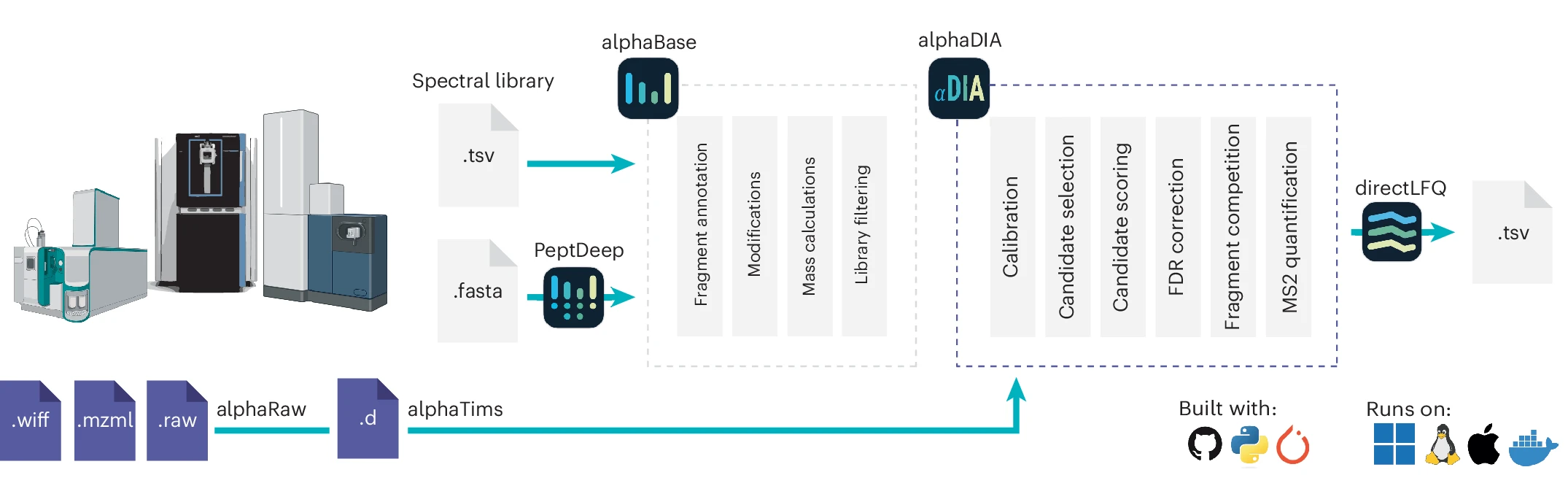
Components of alphaDIA and the integration into the alphaPept ecosystem. AlphaDIA uses alphaRaw and alphaTims for accessing raw data from all major vendors. Importing and prediction of spectral libraries are facilitated by alphaBase and alphapeptdeep. After successful search, LFQ is performed using directLFQ (Figure: Wallmann et al, 2025, Nature Biotech, BioRender).
alphaDIA is fully open, FAIR, and extensible, designed to run locally or in high-performance computing environments, with graphical, command-line, and API access. It follows modern best practices in software engineering — including modular design, continuous integration, automated testing, and rich documentation - to ensure reliability, transparency, and ease of contribution.
Try it yourself!
We warmly invite the community to explore the package, and contribute to alphaDIA on GitHub. Together, we can continue to advance the open, modular, and intelligent future of computational proteomics.
AlphaDIA enables DIA transfer learning for feature-free proteomics Georg Wallmann, Patricia Skowronek, Vincenth Brennsteiner, Mikhail Lebedev, Marvin Thielert, Sophia Steigerwald, Mohamed Kotb, Oscar Despard, Tim Heymann, Xie-Xuan Zhou, Maximilian T. Strauss, Constantin Ammar, Sander Willems, Magnus Schwörer, Wen-Feng Zeng & Matthias Mann, Nature Biotechnology (2025). https://doi.org/10.1038/s41587-025-02791-w